The latest version of this resource was released in August 15 miRSearch It has been shown that bases 27 of 5′ end of the miRNA are crucial to initiate mRNA binding Thus, we suppose that miRNAs, which seed regions are not knotted in secondary structure, interact more easily with target mRNA than those, which seed region is a part of hairpin stem, like eg in the case of miR93 and miR296 The significance of seed region (nucleotides 2–8) thermodynamic stability in the process of offtarget effects has been thoroughly explored 17 Our detailed study revealed that the siRNA seed region encompassing nucleotides 2–5 has the highest positive correlation with offtarget effect ( Fig 2 )

Stability Of Mirna 5 Terminal And Seed Regions Is Correlated With Experimentally Observed Mirna Mediated Silencing Efficacy Topic Of Research Paper In Biological Sciences Download Scholarly Article Pdf And Read For Free On
Seed region of mirna
Seed region of mirna- Here we show that point mutations in the seed region of miR96, a miRNA expressed in hair cells of the inner ear 8, result in autosomal dominant, progressive hearing loss This is the first studyTargetScan is a target prediciton tool that predicts biological targets of miRNAs by searching for the presence of conserved 8mer and 7mer sites that match the seed region of each miRNA The target prediction software is frequently updated;




Got Target Computational Methods For Microrna Target Prediction And Their Extension Abstract Europe Pmc
The nonseed region of siRNA was found to be subdivided into four domains, in which two nucleotide pairs (positions 13 and 14) were replaceable with cognate deoxyribonucleotides without reducing RNAi activity However, Dcr alone can process dsRNA and premiRNA (11, 24–26), but Dcr appears to form a heterodimer with TRBP or PACT Clustered miRNAs with SNPs in seed regions are involved in more pathways, such as Alzheimer disease amyloid secretase pathway, and TCR and IL signaling pathways, whereas nonclustered miRNAs with SNPs in seed regions are only enriched inAbstract MicroRNAs (miRNAs) are studied as key regulators of gene expression involved in different diseases Several single nucleotide polymorphisms (SNPs) in miRNA genes or target sites (miRNArelated SNPs) have been proved to be associated with human diseases by affecting the miRNAmediated regulatory function To systematically analyze miRNArelated SNPs and their effects, we performed a genomewide scan for SNPs in human premiRNAs, miRNA flanking regions
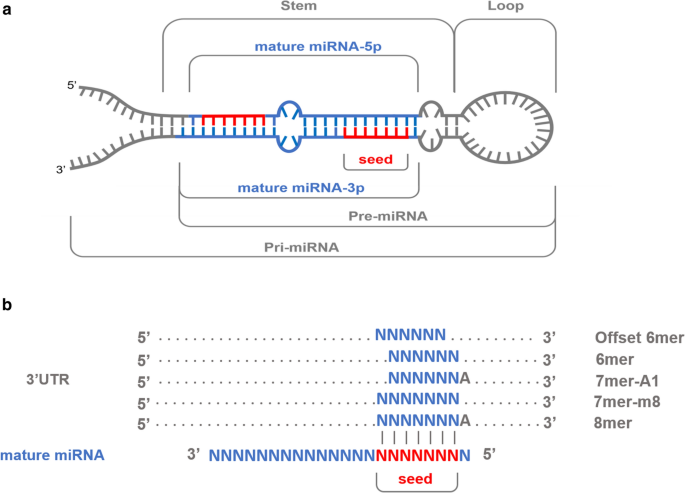
In miRNA targeting, the seed region (nt g2–g7 or g2–g8, from the miRNA 5′ end) is the primary determinant for targeting efficacy and specificity, with over 80% of miRNA–target interactions occurring through seed pairing (Grosswendt et al, 14) MicroRNAs (miRNAs) bind to mRNAs and target them for translational inhibition or transcriptional degradation It is thought that most miRNAmRNA interactions involve the seed region at the 5′ end of the miRNA The importance of seed sites is supported by experimental evidence, although there is growing interest in interactions mediated by the central region of the miRNASense strand), also called the 'seed region', is complementary to the 3′ untranslated regions (UTRs) of multiple mRNAs, causing degradation of their associated transcripts 8,9 To improve the interpretation of RNAi datasets and to help minimize followup experimental efforts, it is important to identify transcripts that are
Most miRNAs pair to the target sites through seed region near the 5' end, leading to mRNA cleavage and/or translation repression Here, we demonstrated a miRNAinduced dual regulation of heme oxygenase1 (HO1) via seed region and nonseed region, consequently inhibited tumor growth of NSCLC miRNA seed region enrichment in the exosomal lncRNA is independent of sequence length To investigate if sequence length was a determinant in miRNA seed enrichment, we compared the average lengthSpecifically, seedbased canonical target recognition was dependent on the GC content of the miRNA seed For miRNAs with low GC content of the seed region, noncanonical targeting was the dominant mechanism for target recognition
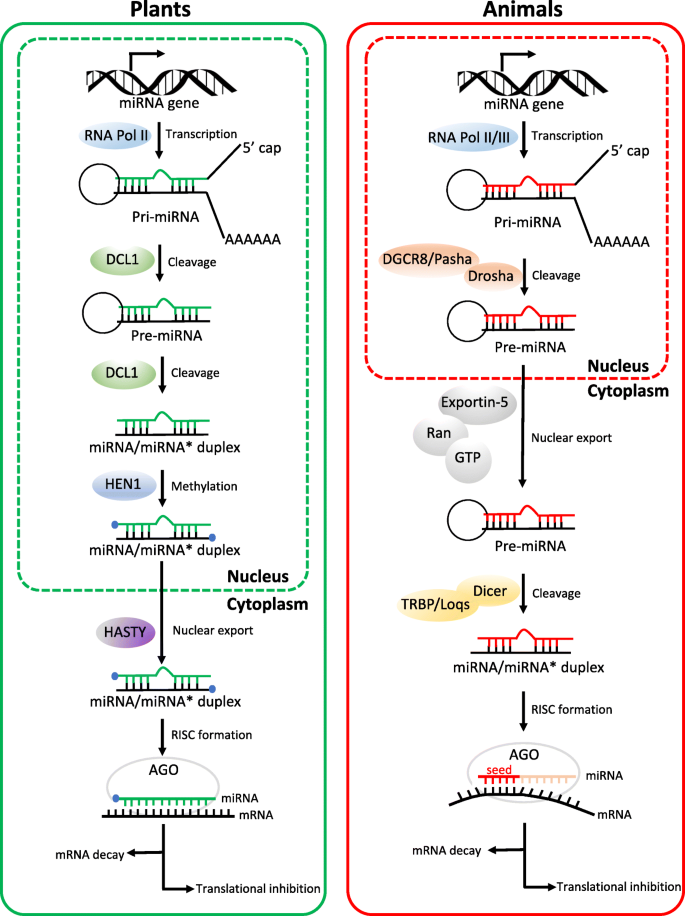



Micrornas From Plants To Animals Do They Define A New Messenger For Communication Nutrition Metabolism Full Text




Example Of Mirna Target Alignment This Schematic Representation Shows Download Scientific Diagram
Among all seedpairing target sites (9465 in total), 435% of them were also paired to nonseed region of miRNA, which was a significant enrichment over shuffled control (Table 2) Furthermore, as to noncanonical target sites without any seed match, 409% were paired to the nonseed region of the miRNAAnd some miRNA nucleotides are more important than others (8, 9) Specifically, pairing to the miRNA "seed region" (nt 2 to 7 or 2 to 8, from the 5′ end) is the most evolutionarily conserved feature of miRNA targets in animals (10–14) Crystal structures of human Argonaute proteins show nt 2 to 6 of the guide RNA bound A mismatch tolerance test assay, based on pools of transgenic strains, revealed that target hybridization to nucleotides of the seed region, at the 5′ end of an miRNA, was sufficient to induce moderate repression of expression In contrast, pairing to the 3′ region of the miRNA was not critical for silencing



Targetscan Non Canonical Sites
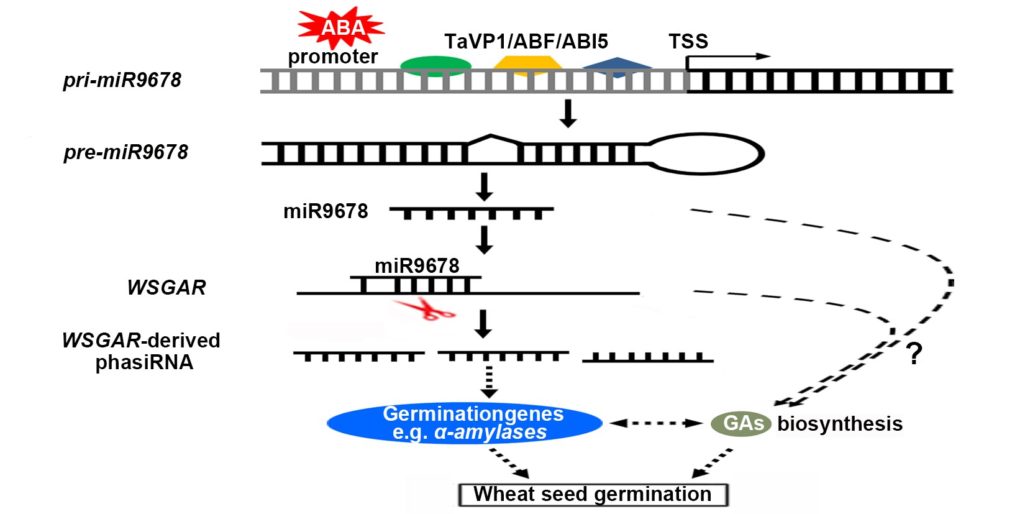



Plantae Mirna Mediated Regulation Of Germination Plantae
The miRNA sequence can be separated into five functional domains that affect miRNAtarget recognition 5′ anchor (nt 1), seed sequence (nts 2–8), central region (nts 9–12), 3′ supplementary region (nts 13–16), and 3′ tail (nts 17–22) (Wee et al, 12) We anticipated that complementarity to the seed sequence of the cognate miRNA would be a prominent feature inIn human miRNA seed regions A total of 1879 SNPs were mapped to 1226 human miRNA seed regions We found that miRNAs with SNPs in their seed region are significantly enriched in miRNA clusters We also found that SNPs in clustered miRNA seed regions have a lower functional effect than have SNPs in nonclustered miRNA seed regions Additionally, we This socalled seed region is the largest contributor in determining miRNA target specificity (reviewed in Ref 73) As such, changing the 5′ terminus leads to a shift in the seed region and can




Dbmts A Comprehensive Database Of Putative Human Microrna Target Site Snvs And Their Functional Predictions Li Human Mutation Wiley Online Library
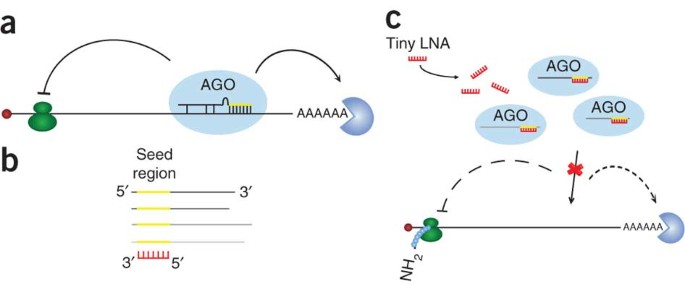



Silencing Of Microrna Families By Seed Targeting Tiny Lnas Nature Genetics
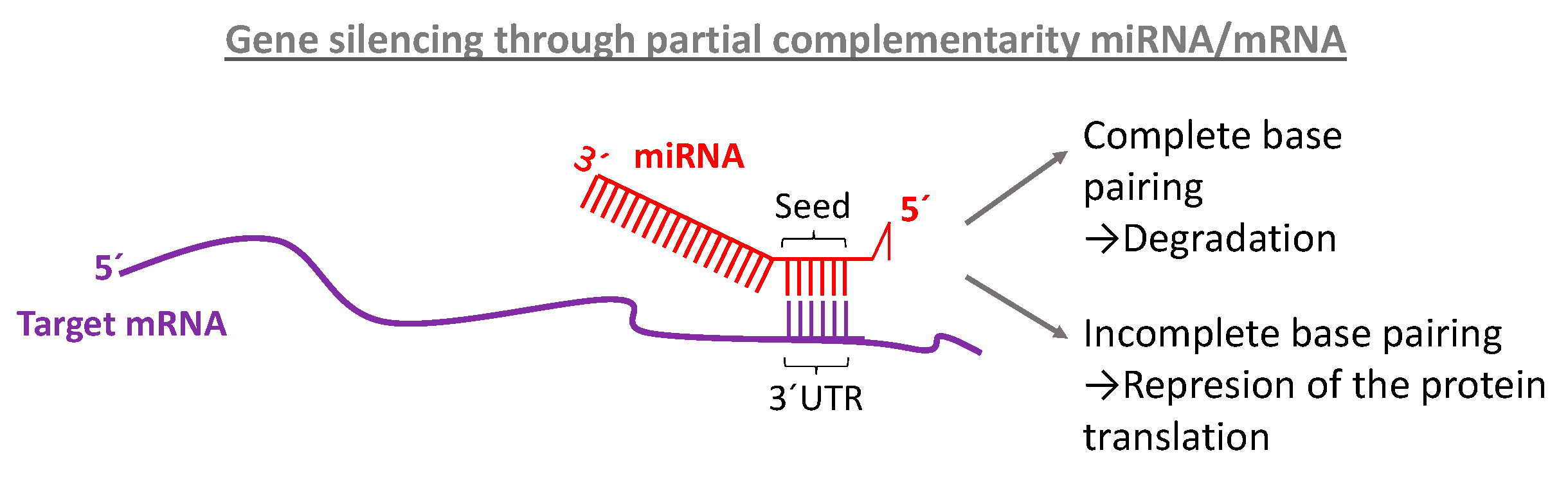
The most conserved part of miRNA is located at 5′end and is called the seed region Guide (g) nucleotides from the seed region (g2–g7 or g2–g8) play a primary role in target recognition because more than 80% of the interactions between miRNAs and targets occurs via seed The apparent importance of seed pairing determined by computational studies was perfectly explained by later structural insights miRNA forms a complex with Ago in a unique structural organization where only the seed region is exposed and available to pair with target mRNAs The first nucleotide of miRNA is inserted inside the Ago protein and does notMiRNA This interaction between the miRNA and 39 UTR is dependent upon a 6–8 nucleotide sequence found in the 59 end of the miRNA called the ''seed'' sequence This sequence must have perfect complementarity with its 39 UTR binding site for repression to occur Disruption of the seed region of the miRNA




Rna Mrna Target Interaction Schematic Overview Of A Mirna Interaction Download Scientific Diagram
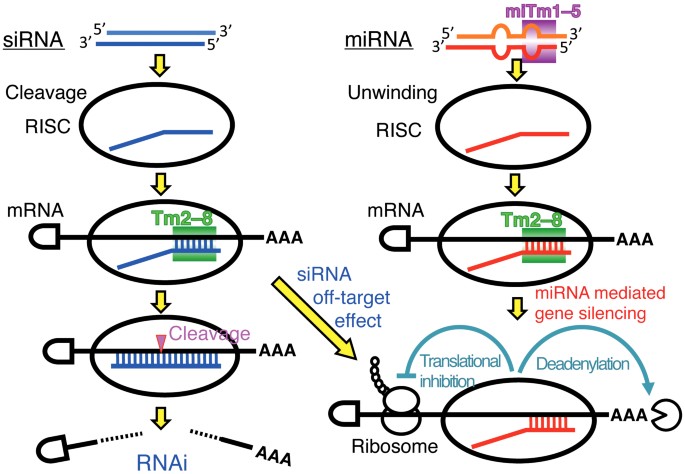



Stability Of Mirna 5 Terminal And Seed Regions Is Correlated With Experimentally Observed Mirna Mediated Silencing Efficacy Scientific Reports
Predicts biological targets of miRNAs by searching for the presence of sites that match the seed region of each miRNA In flies and nematodes, predictions are ranked based on the probability of their evolutionary conservation In zebrafish, predictions are ranked based on site number, site type, and site context, which includes factors that Base pairing of the miRNA seed region (positions 2–8 from the 5′ end of an miRNA) in animals is critical for target recognition and repression (Bartel, 09;Seed region of miRNA associated with Argonaute2 protein The seed region of the miRNA strand is shown Nucleotides 2 through 5 can be seen in positions that are able to interact with the complementary mRNA strand Nucleotides 1, 6 and 7 are not exposed to the mRNA stand in this conformation Seed region of miRNA




A Cartoon Showing The Site And Mechanism Of Mirna Targeting To Mrna Download Scientific Diagram




Human Polymorphism At Micrornas And Microrna Target Sites Pnas
Moreover, the requirement of a 7nt match to the seed region of the miRNA (nucleotides 2–8) could be relaxed to require a 6nt match to a reduced seed comprising nucleotides 2–7 of the miRNA while still retaining modest specificity Running TargetScan in this way without cutoffs amounted to predicting a target simply by virtue of theMarin and Vanicek 11) Evidence from crystal structures, however, suggests that the 5′end of the seed match—more exactly positions 7 b and 8 b (notation defined in Fig 4A )—might not Design of miRNA seedtargeting tiny LNAs Our goal was to develop an antimiR approach that would enable inhibition of miRNA families by targeting their shared seed region ()We have previously




Prediction Of The Mirna Interactome Established Methods And Upcoming Perspectives Sciencedirect
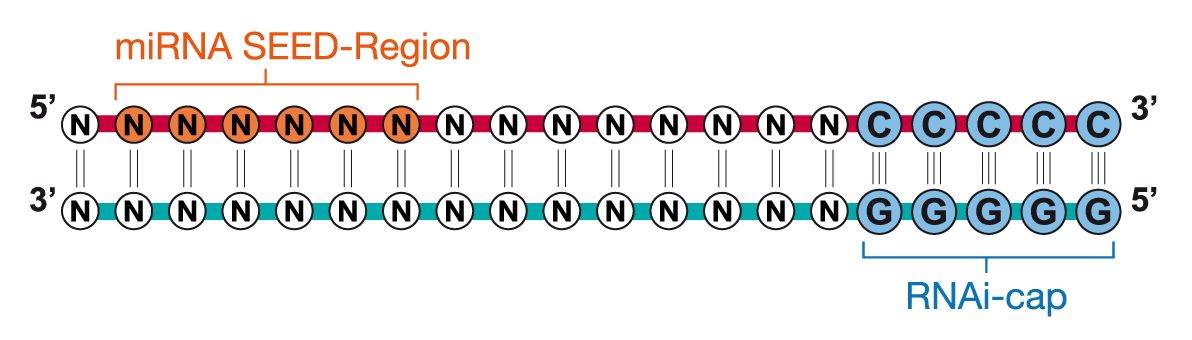



Riboxx Rna Technologies Benefits Of Rnai Cap For Mirna
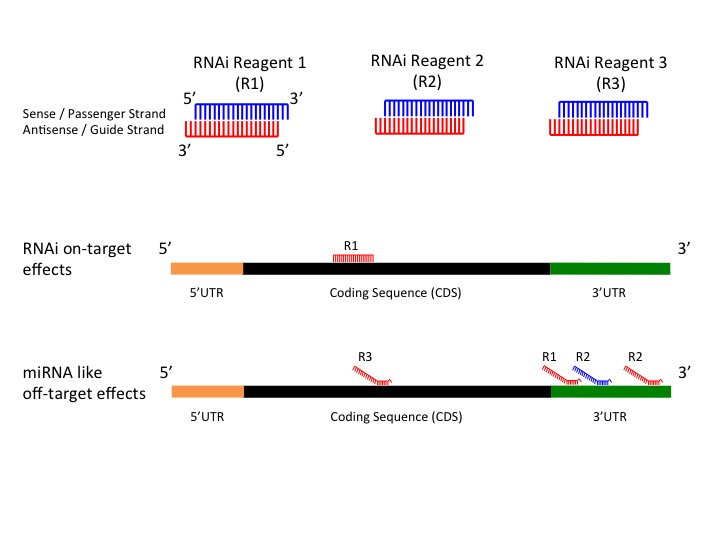
SOFTWARE Open Access Online GESS prediction of miRNAlike offtarget effects in largescale RNAi screen data by seed region analysis Bahar Yilmazel1, Yanhui Hu1, Frederic Sigoillot2, Jennifer A Smith3, Caroline E Shamu3, Norbert Perrimon1,4 and Stephanie E Mohr1* Abstract One class of pattern consists of perfect WatsonCrick binding at the 5'end of the miRNA This region is known as "seedregion" and found at the 27 base of the miRNA This region is able to suppress the target mRNAs without having a complete base pairing atOutside of the seed region, 2 variants in mature regions, 5 in arm regions, 1 in the loop, and 19 within flanking regions of miRNAs also had raw pvalues less than 005 None of these variants were predicted to have an impact on miRNA seed region by all three algorithms implemented in miRVaS ( Table S2 )



2



Mir0c Microrna 0c
Tematic mutagenesis studies of miRNA/target site hybrids have revealed that themiRNA seed region is more critical than the 3′ region fortarget recognition in A thaliana (Mallory et al, 04) Moreover, plant and algal small RNAs also induce translationalrepression ofPasquinelli, 12) In contrast, most evidence indicates that miRNAs in land plants require more extensive pairing to their targets (Schwab et al , 05 ;MicroRNA editing in seed region aligns with cellular changes in hypoxic conditions RNA editing is a finely tuned, dynamic mechanism for posttranscriptional gene regulation that has been thoroughly investigated in the last decade




Stability Of Mirna 5 Terminal And Seed Regions Is Correlated With Experimentally Observed Mirna Mediated Silencing Efficacy Topic Of Research Paper In Biological Sciences Download Scholarly Article Pdf And Read For Free On



2
Several miRNAs have been shown to play an essential role in adipogenesis, such as miR27 16, miR21 17, miR27b 18, and miR335 19 The SNP identified in our study is located in the seed region of the miRNA that has to be complementary to the mRNA in The distribution pattern of genetic variation in miRNA seed regions might be related to miRNA function and is worth paying more attention to We here employed computational analyses to explore the clustering pattern and functional effect of SNPs in human miRNA seed regions A total of 1879 SNPs were mapped to 1226 human miRNA seed regionsMiRNAs regulate the gene expression by binding to the mRNA The seed sequence is essential for the binding of the miRNA to the mRNA The seed sequence or seed region is a conserved heptametrical sequence which is mostly situated at positions 27 from the miRNA 5´end Even though base pairing of miRNA and its target mRNA does not match perfect, the "seed sequence"
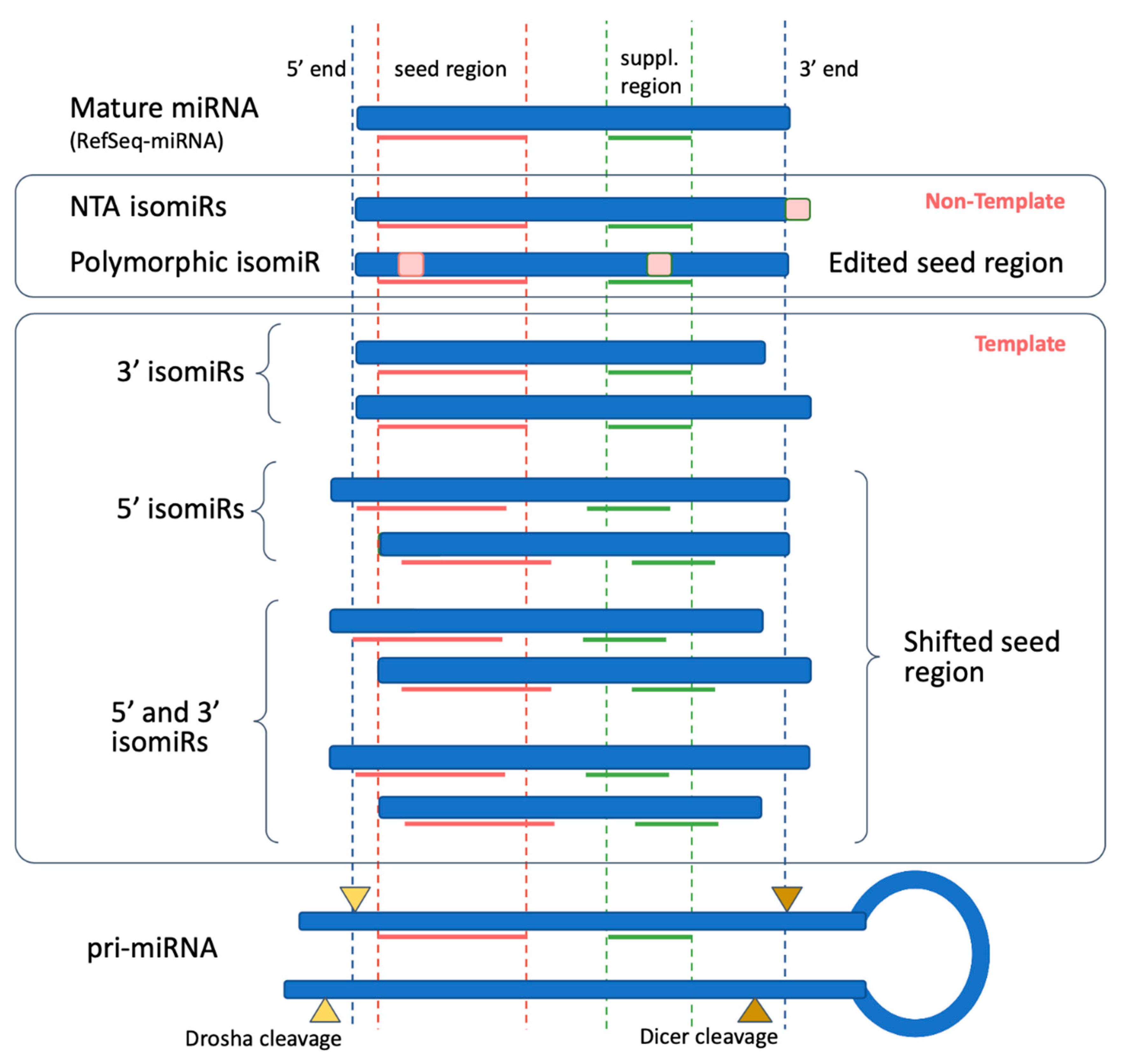



Biomolecules Free Full Text Isomirs Hidden Soldiers In The Mirna Regulatory Army And How To Find Them Html




Function Control Of Anti Microrna Oligonucleotides Using Interstrand Cross Linked Duplexes Molecular Therapy Nucleic Acids
MicroRNAs, or miRNAs, posttranscriptionally repress the expression of proteincoding genes The human genome encodes over 1000 miRNA genes that collectively target the vast majority of messenger RNAs (mRNAs) Basepairing of the socalled miRNA "seed" region with mRNAs identifies many thousands of putative targetsComputational methods play an important role in target prediction and assume that the miRNA "seed region" (nt 2 to 8) is required for funct MicroRNAs (miRNAs) are small noncoding RNAs that regulate gene expression by binding to partially complementary regions within the 3'UTR of their target genes Computational methods play an important role in target prediction and assume that the miRNA "seed region" Thus, the functions of miRNA 5′ terminal and seed regions in miRNAmediated gene silencing may differ Some miRNAs, such as let7 47,48, miR34 49,




Got Target Computational Methods For Microrna Target Prediction And Their Extension Abstract Europe Pmc




Micro Arn Wikipedia La Enciclopedia Libre
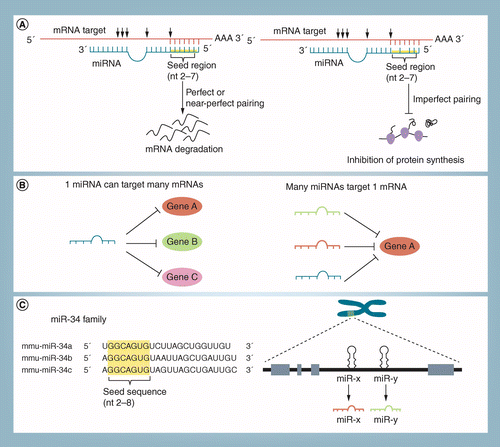
These molecules typically reduce the translation and stability of mRNAs, including those of genes that mediate processes in tumorigenesis, such as inflammation, cell cycle regulation, stress response, differentiation, apoptosis, and invasion miRNA targeting is initiated through specific basepairing interactions between the 5' end ("seed" region) of the miRNA and sites within The miRNA target prediction methods requiring a nucleation region usually allow this region to be located anywhere within the seed (Robins et al 05;MicroRNA (miRNA), originally discovered in Caenorhabditis elegans, is found in most eukaryotes, including humans 13 All four miRNAs share a 9nucleotide seed region that targets the 3'UTR of the antiapoptotic protein BCL2 for posttranscriptional repression 261, 263




Microrna 145 Modulates N6 Methyladenosine Levels By Targeting The 3 Untranslated Mrna Region Of The N6 Methyladenosine Binding Yth Domain Family 2 Protein Journal Of Biological Chemistry



2



Plos One Functional And Evolutionary Significance Of Human Microrna Seed Region Mutations



2



Plos One Evolution Of Microrna In Primates




Mirna Mirna Targets Functional Analysis



1




Crispr Screening Strategies For Microrna Target Identification Yang The Febs Journal Wiley Online Library
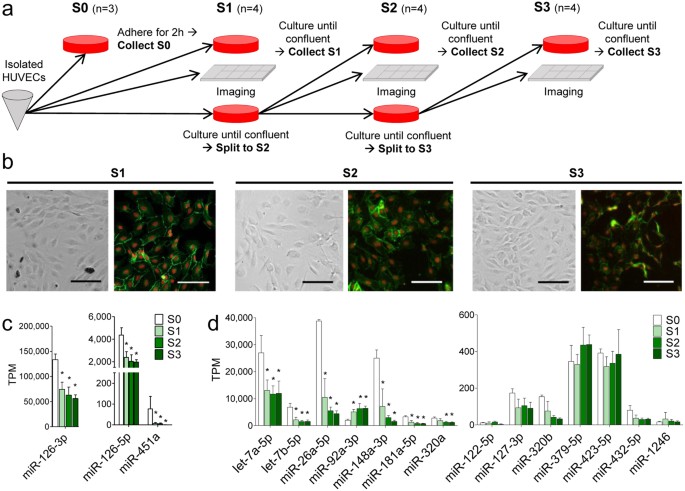



Microrna Profiling Reveals Distinct Profiles For Tissue Derived And Cultured Endothelial Cells Scientific Reports
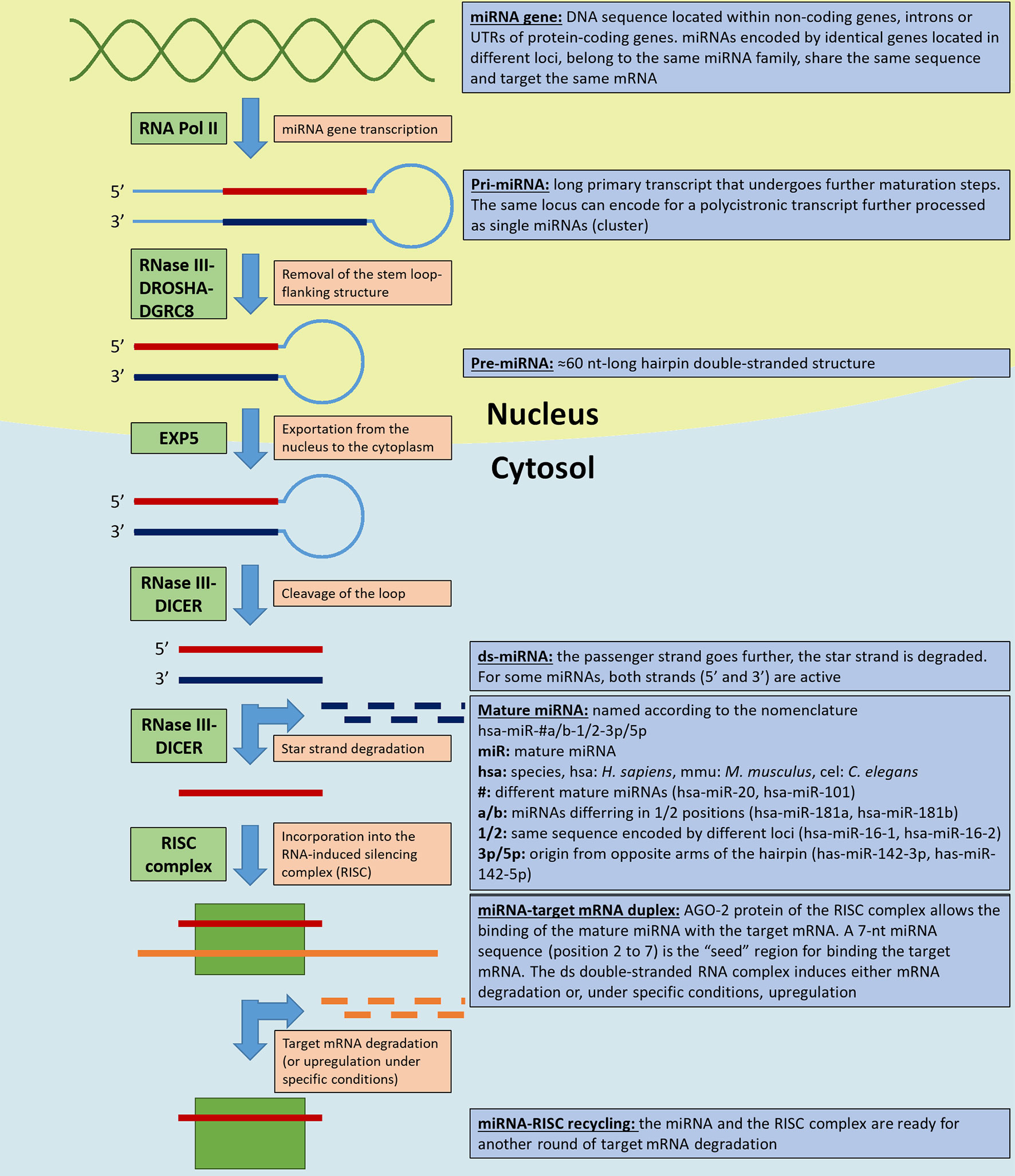



Frontiers Perspectives On Mirnas As Epigenetic Markers In Osteoporosis And Bone Fracture Risk A Step Forward In Personalized Diagnosis Genetics



Mirvestigator Framework Detect The Mirnas Driving Co Expression Signatures



Gess




Mirna Seed Region Shefalitayal




Beyond The Seed Structural Basis For Supplementary Microrna Targeting By Human Argonaute2 The Embo Journal




Mapping The Human Mirna Interactome By Clash Reveals Frequent Noncanonical Binding Abstract Europe Pmc




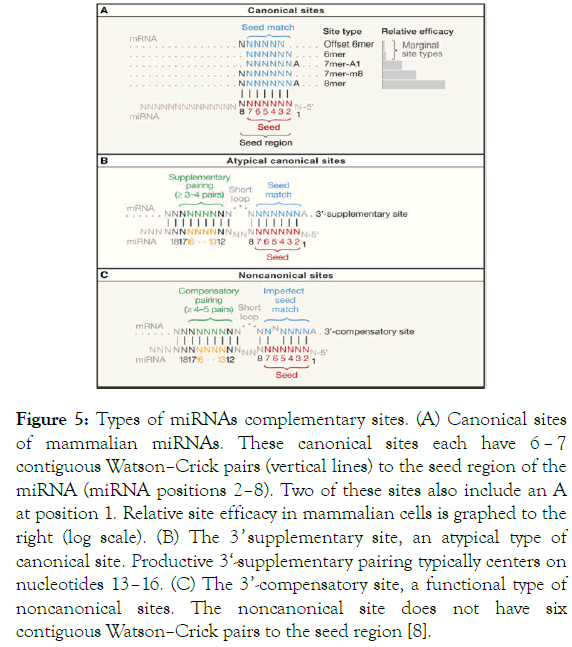
Types Of Microrna Target Sites A There Are Three Types Of Canonical Download Scientific Diagram
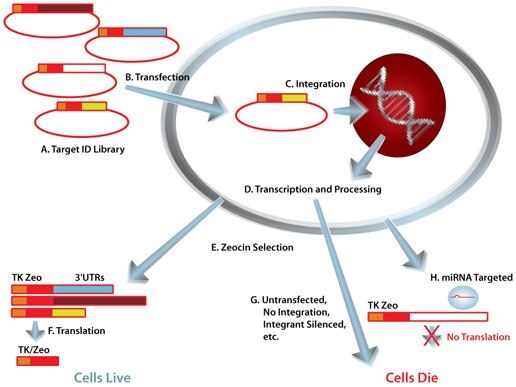



Mission Target Id Library For Human Mirna Target Id And Discovery




Chemical Modifications In The Seed Region Of Mirnas 221 222 Increase The Silencing Performances In Gastrointestinal Stromal Tumor Cells Sciencedirect



1
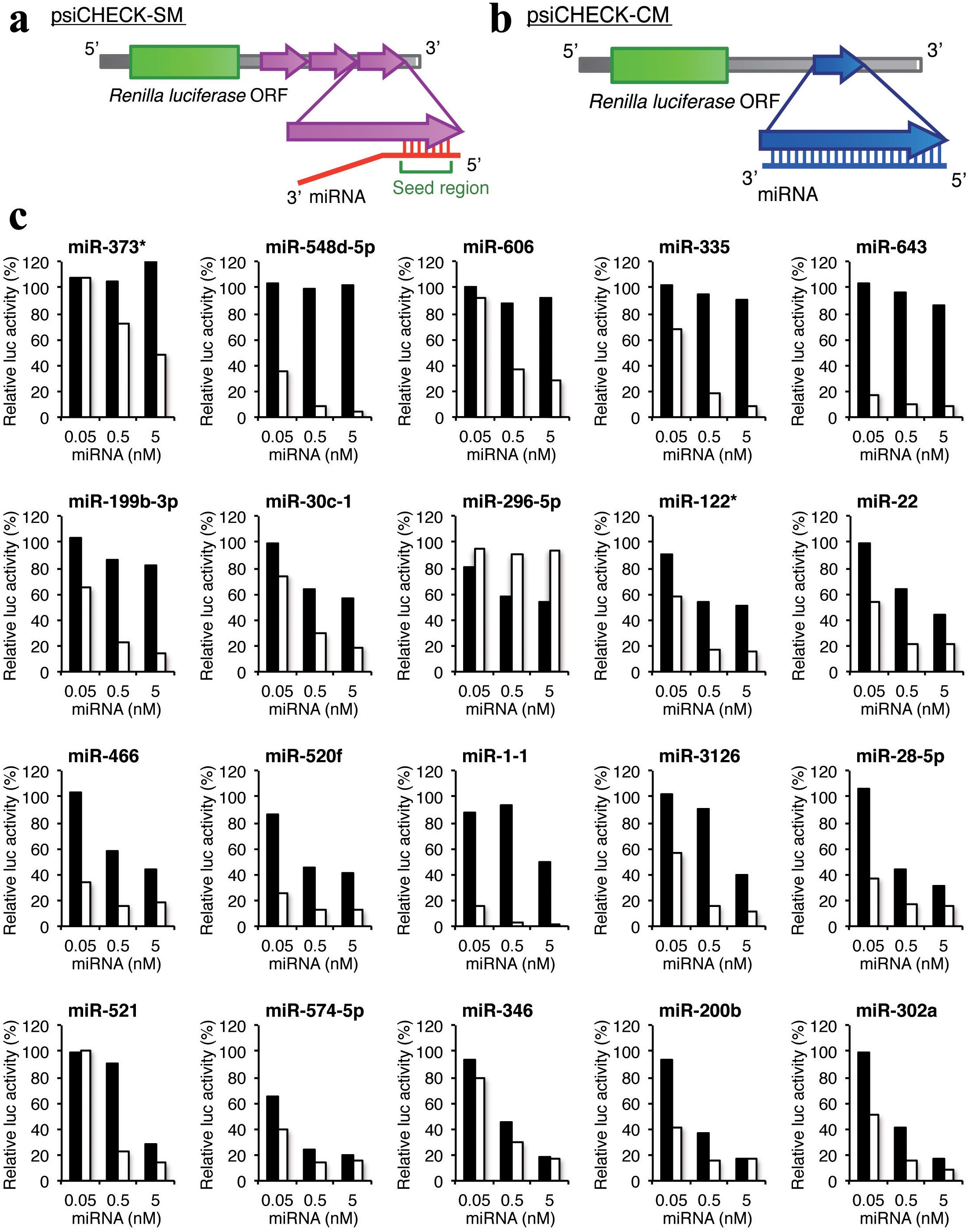



Stability Of Mirna 5 Terminal And Seed Regions Is Correlated With Experimentally Observed Mirna Mediated Silencing Efficacy Scientific Reports




Microrna Regulation And Cardiac Calcium Signaling Circulation Research



Micrornas In Skin Biology Biogenesis Regulations And Functions In Homeostasis And Diseases




Mirna Targeting Growing Beyond The Seed Trends In Genetics




The Biochemical Basis Of Microrna Targeting Efficacy




Microrna Wikipedia




Eshre19




Microrna Gga Mir 6578 Containing Polymorphism Within The Mirna Seed Download Scientific Diagram




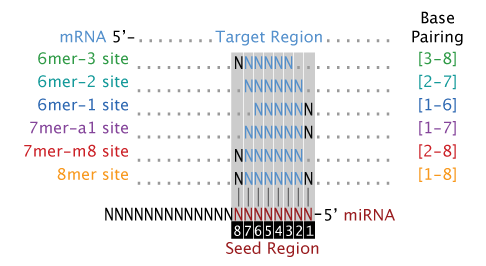
Different Mirna Mrna Seed Site Interaction Patterns 6mer 7mer A1 Download Scientific Diagram




Mirna Introduction Biogenesis Nomenclature And Experimental Workflow



Plos One Micrornas Mir 19 Mir 340 Mir 374 And Mir 542 Regulate Mid1 Protein Expression




Miraw A Deep Learning Approach To Predict Mirna Targets By Analyzing Whole Mirna Transcripts Biorxiv




Target Recognition By Sirna And Mirna A Sirna Is Usually Fully Download Scientific Diagram
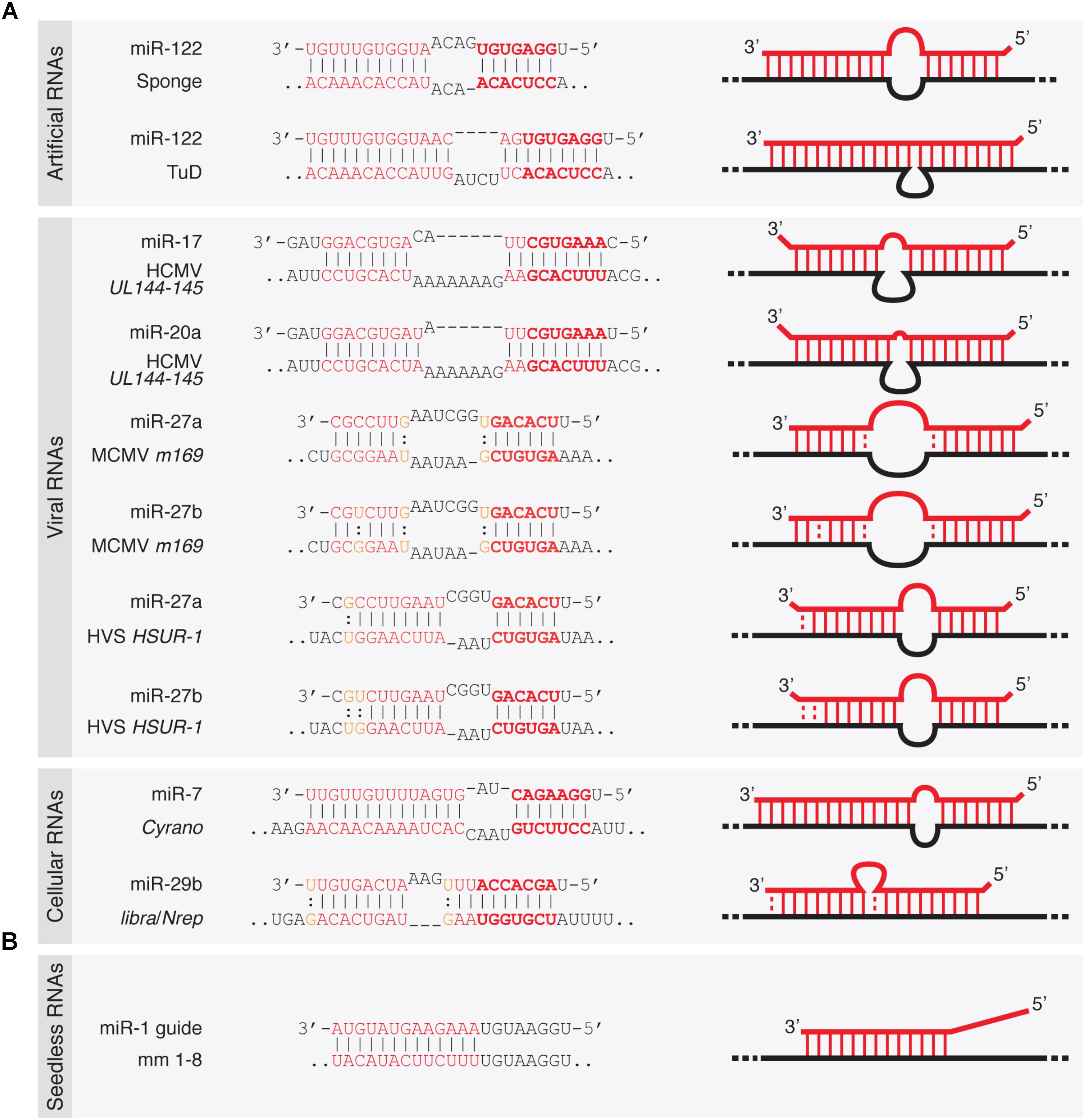



Frontiers Target Rnas Strike Back On Micrornas Genetics




Regulatory Network Of Mirna On Its Target Coordination Between Transcriptional And Post Transcriptional Regulation Of Gene Expression Semantic Scholar




Helix 7 In Argonaute2 Shapes The Microrna Seed Region For Rapid Target Recognition The Embo Journal




Miraw A Deep Learning Approach To Predict Mirna Targets By Analyzing Whole Mirna Transcripts Biorxiv
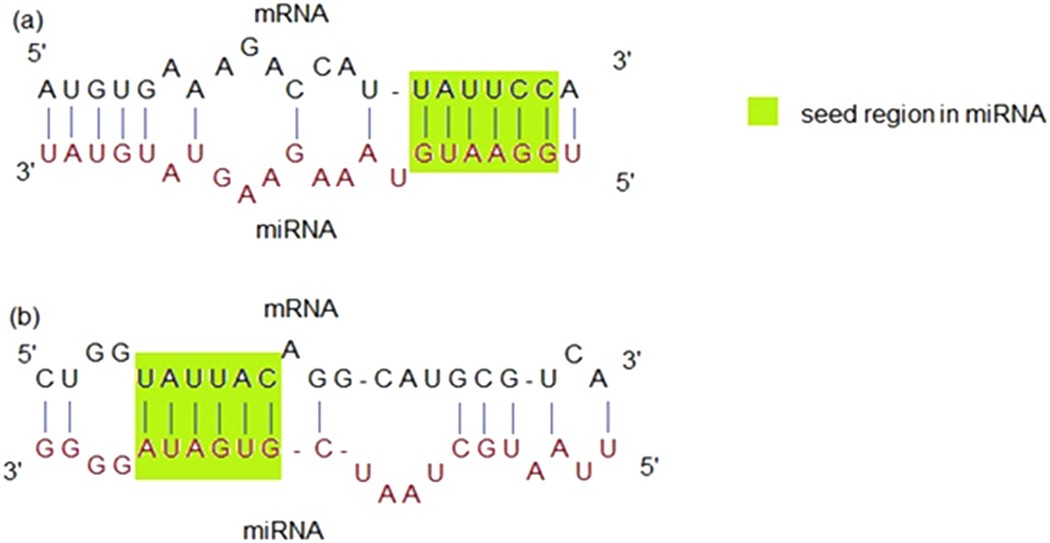



Mirepress Modelling Gene Expression Regulation By Microrna With Non Conventional Binding Sites Scientific Reports




Processes Free Full Text Micrornas Biogenesis Functions And Potential Biomarkers For Early Screening Prognosis And Therapeutic Molecular Monitoring Of Nasopharyngeal Carcinoma Html
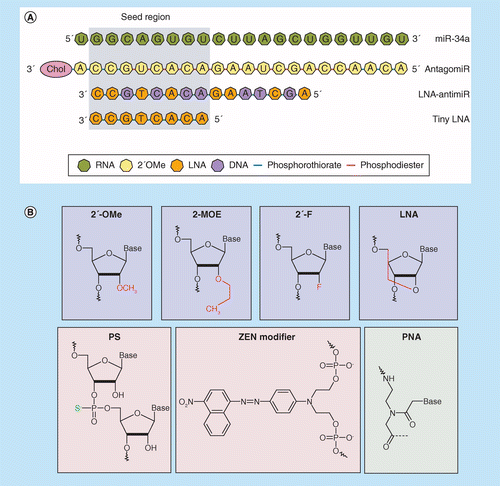



Mirna Therapeutics A New Class Of Drugs With Potential Therapeutic Applications In The Heart Future Medicinal Chemistry
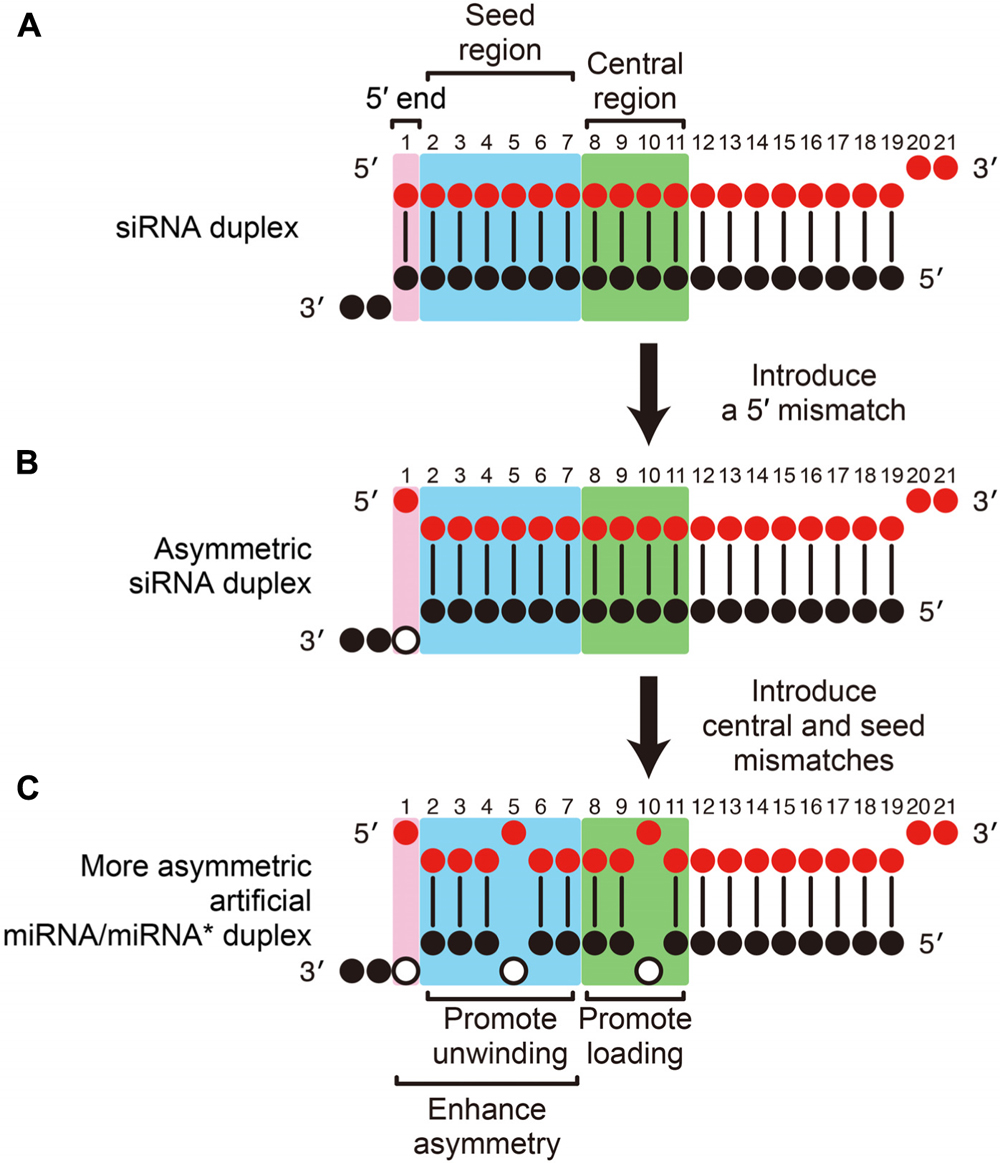



Frontiers Mirna Like Duplexes As Rnai Triggers With Improved Specificity Genetics




The Characterization Of Microrna Mediated Gene Regulation As Impacted By Both Target Site Location And Seed Match Type




Mirna 27b Targets The 3 Utr Of Foxj3 A Sequence Alignment Of Download Scientific Diagram




Mismatches In The Mirna Proximal Seed Region Disrupt The Binding Download Scientific Diagram




Mirna Introduction Biogenesis Nomenclature And Experimental Workflow




Examples Of Mirna Target Interactions Pairing Schemes Of Several Download Scientific Diagram




Microrna Polymorphisms The Future Of Pharmacogenomics Molecular Epidemiology And Individualized Medicine Pharmacogenomics
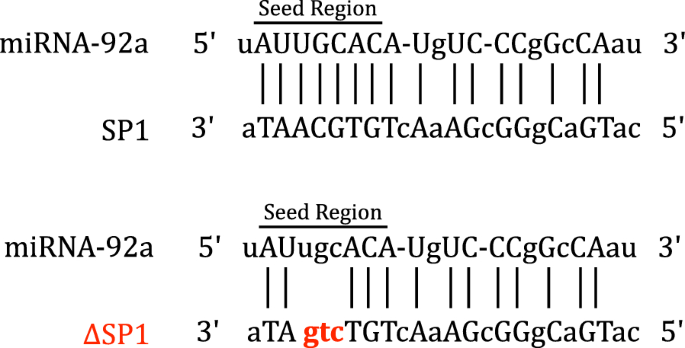



Microrna 92a Regulates The Expression Of Aphid Bacteriocyte Specific Secreted Protein 1 Bmc Research Notes Full Text
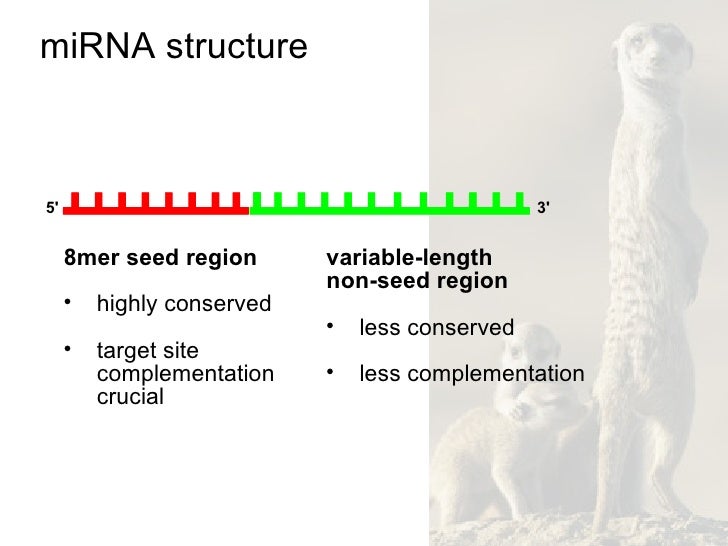



Human Snps In Microrna Target Sites
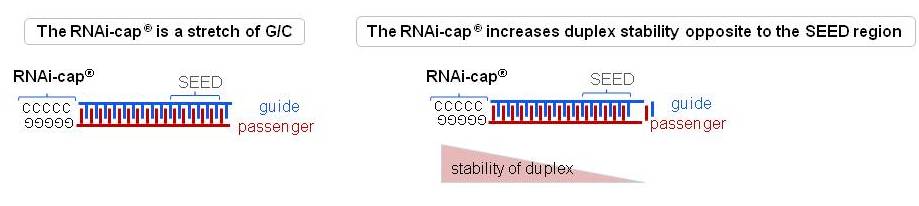




Riboxx Rna Technologies Benefits Of Rnai Cap For Mirna
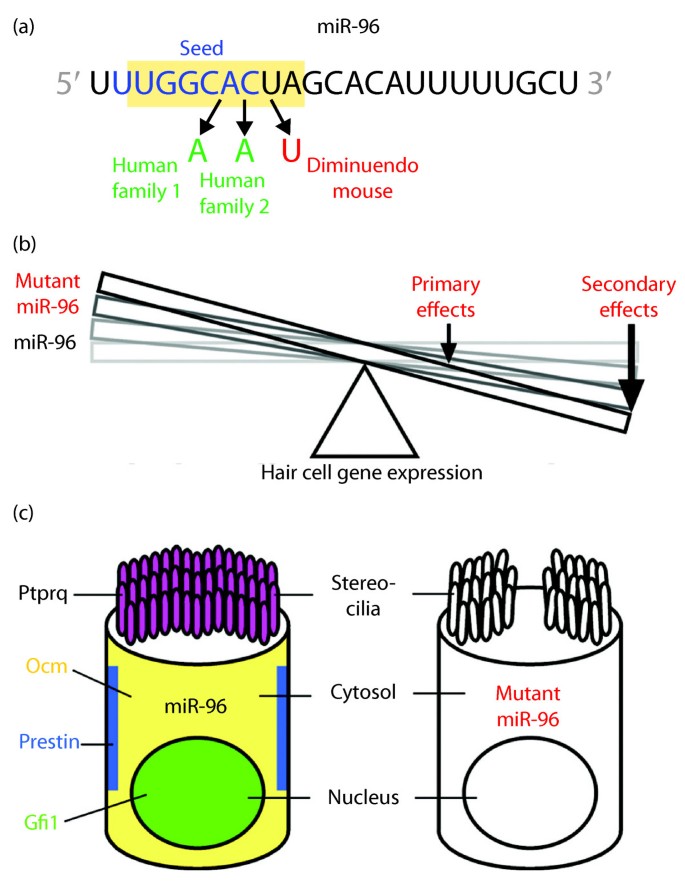



Micrornas Sound Off Genome Medicine Full Text



Ijms Free Full Text Microrna Nanotherapeutics For Lung Targeting Insights Into Pulmonary Hypertension Html



Variability In Porcine Microrna Genes And Its Association With Mrna Expression And Lipid Phenotypes Genetics Selection Evolution Full Text



Endogenous Transcripts Control Mirna Levels And Activity In Mammalian Cells By Target Directed Mirna Degradation Nature Communications




A Study Of Micrornas In Silico And In Vivo Bioimaging Of Microrna Biogenesis And Regulation Kim 09 The Febs Journal Wiley Online Library



1




Pairing Beyond The Seed Supports Microrna Targeting Specificity Sciencedirect




Mapping The Human Mirna Interactome By Clash Reveals Frequent Noncanonical Binding Cell




6mer Seed Toxicity In Viral Micrornas Sciencedirect




Mapping The Human Mirna Interactome By Clash Reveals Frequent Noncanonical Binding Sciencedirect




Correlation Between Mirna Targeted Gene Promoter F1000research




Mirna Seed Types Nine Seed Types Are Categorized In Two Groups Download Scientific Diagram




Beyond The Seed Structural Basis For Supplementary Microrna Targeting By Human Argonaute2 The Embo Journal



1



Mirna Therapeutics A New Class Of Drugs With Potential Therapeutic Applications In The Heart Future Medicinal Chemistry



Microarn Viquipedia L Enciclopedia Lliure
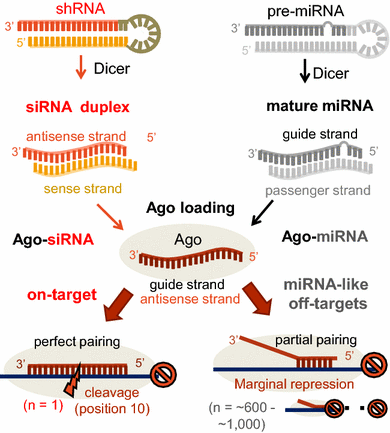



Evaluation And Control Of Mirna Like Off Target Repression For Rna Interference Springerlink




Different Seed Match Regions Of Mirnas Mirnalyze Follows A Download Scientific Diagram




Sites Matching In The Mirna Seed Region Including All K Mer 8mer Download Scientific Diagram




The Biochemical Basis Of Microrna Targeting Efficacy Biorxiv




Extracellular Micrornas In Human Circulation Are Associated With Mirisc Complexes That Are Accessible To Anti Ago2 Antibody And Can Bind Target Mimic Oligonucleotides Pnas




Types Of Mirna Target Sites And Multiple Sites A Stringent Seed Download Scientific Diagram




6mer Seed Toxicity In Viral Micrornas Biorxiv




Beyond The Seed Structural Basis For Supplementary Microrna Targeting By Human Argonaute2 The Embo Journal




Pairing Beyond The Seed Supports Microrna Targeting Specificity Sciencedirect




Dbmts A Comprehensive Database Of Putative Human Microrna Target Site Snvs And Their Functional Predictions Li Human Mutation Wiley Online Library



2




Systematic Prediction Of The Impacts Of Mutations In Microrna Seed Sequences




6mer Seed Toxicity In Viral Micrornas Biorxiv




Mirna Therapeutics A New Class Of Drugs With Potential Therapeutic Applications In The Heart Future Medicinal Chemistry
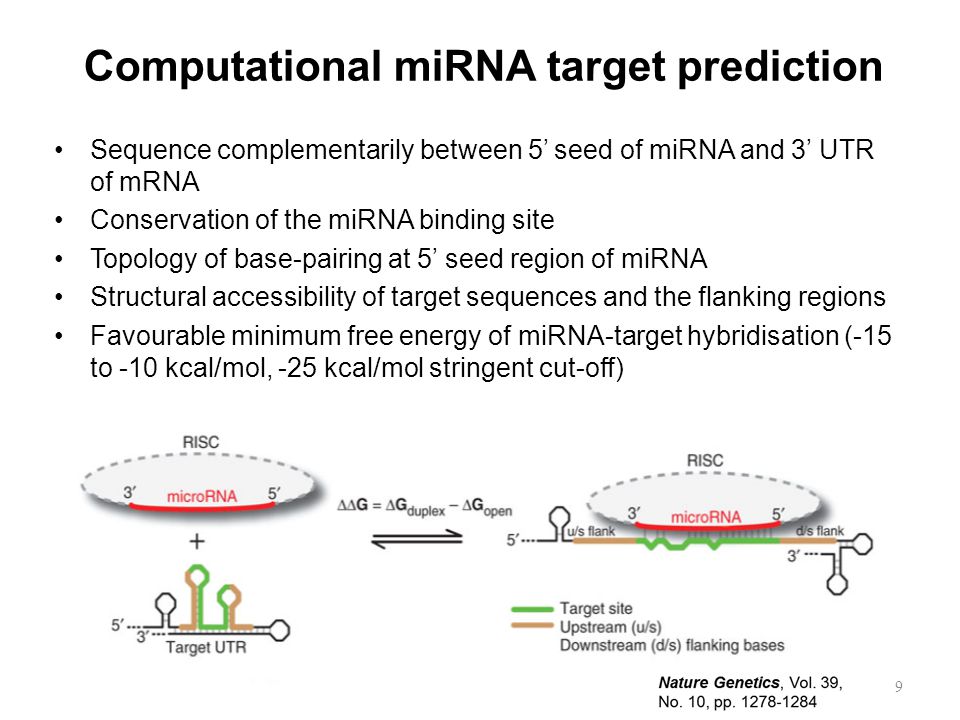



Methods For Identifying Microrna Binding Motifs Ppt Video Online Download



Plos One Microrna Seed Region Length Impact On Target Messenger Rna Expression And Survival In Colorectal Cancer
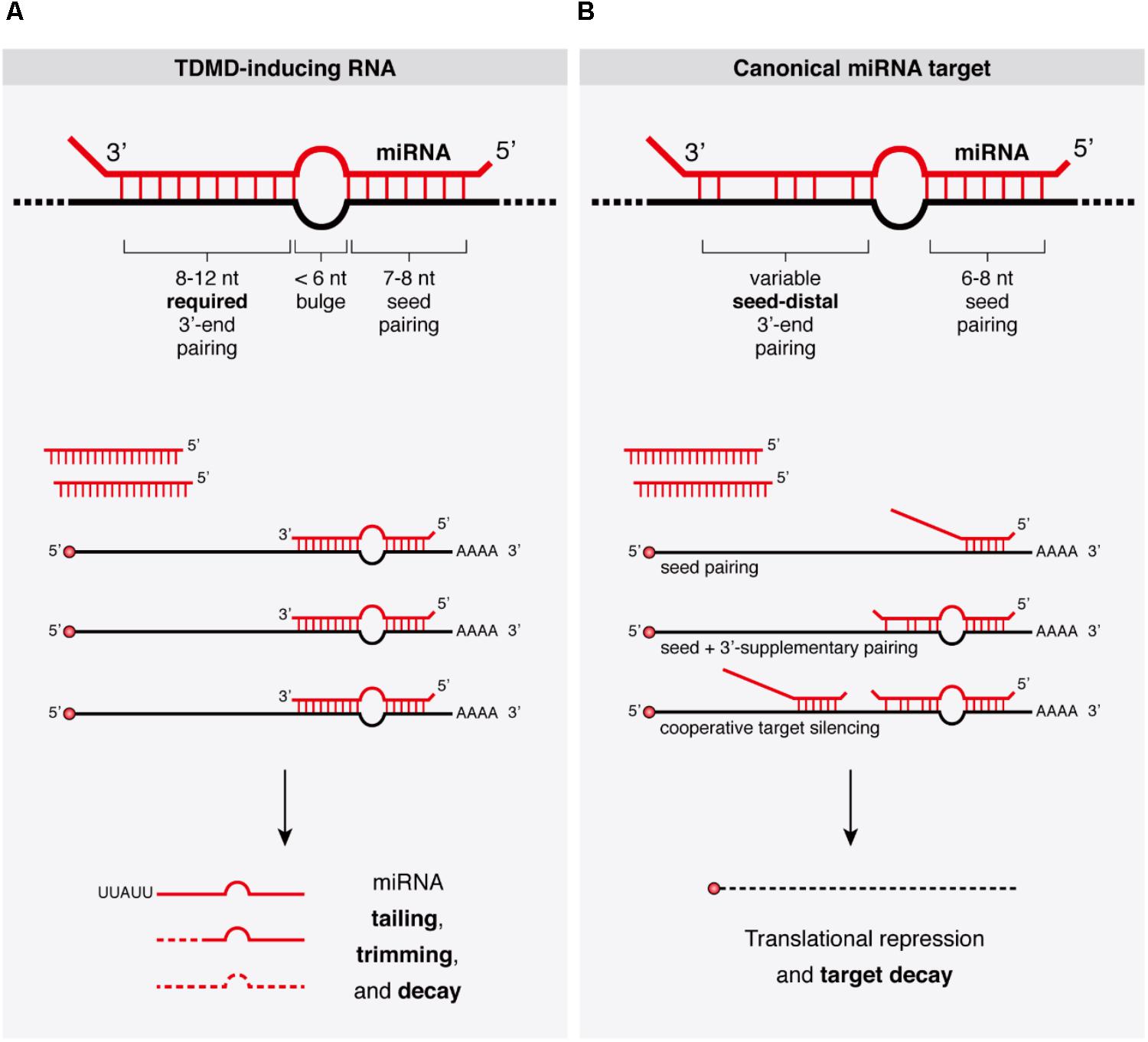



Frontiers Target Rnas Strike Back On Micrornas Genetics




Mirna Sequencing Report



2



0 件のコメント:
コメントを投稿